
Calculate the probability of metabolite formation and screen for toxicity potential
META Ultra uses a data-driven approach to analyze the atomic environment around each site of metabolism (SOM) up to a depth of 3 bonds.
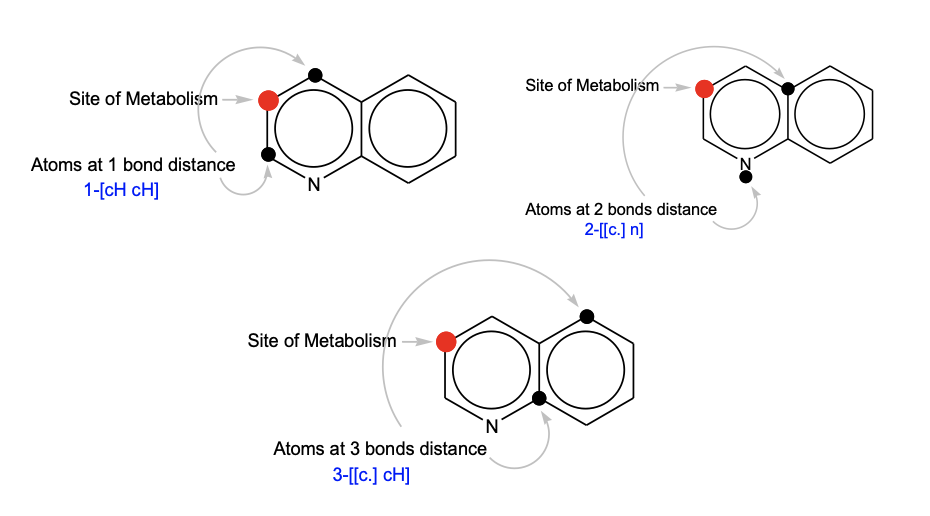
The probability of the formation of Phase I and Phase II metabolites are predicted using (Q)SAR.
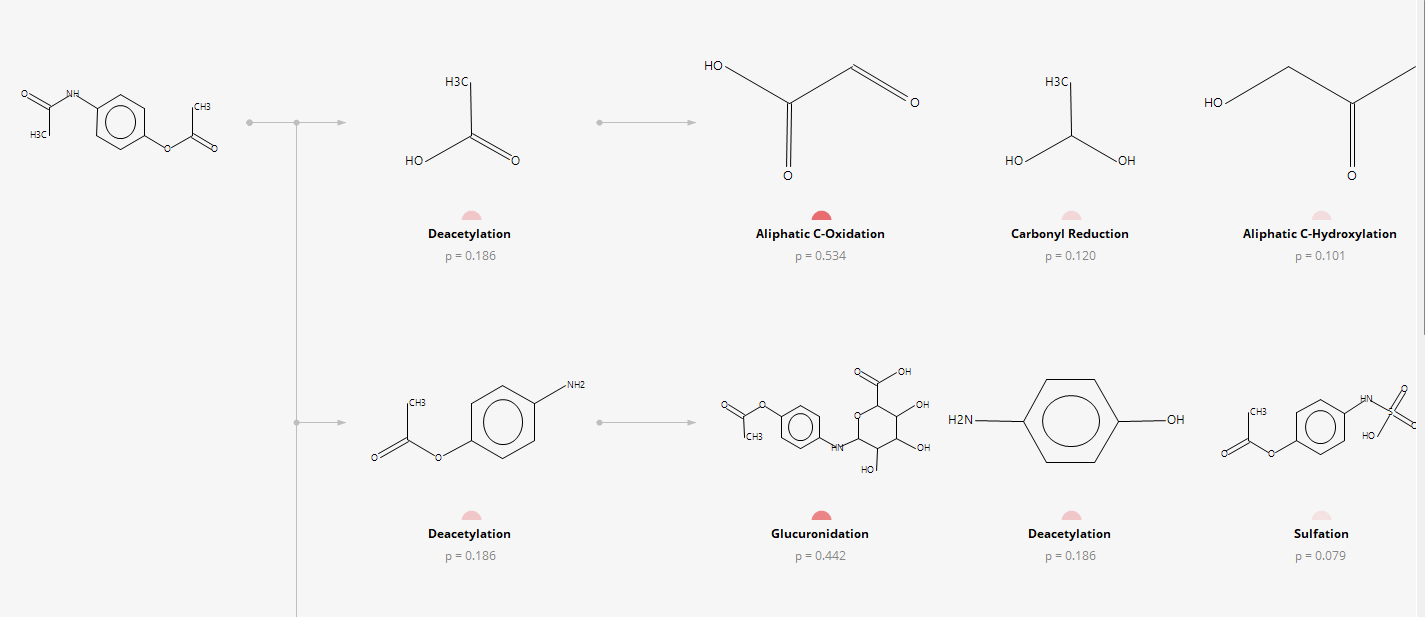
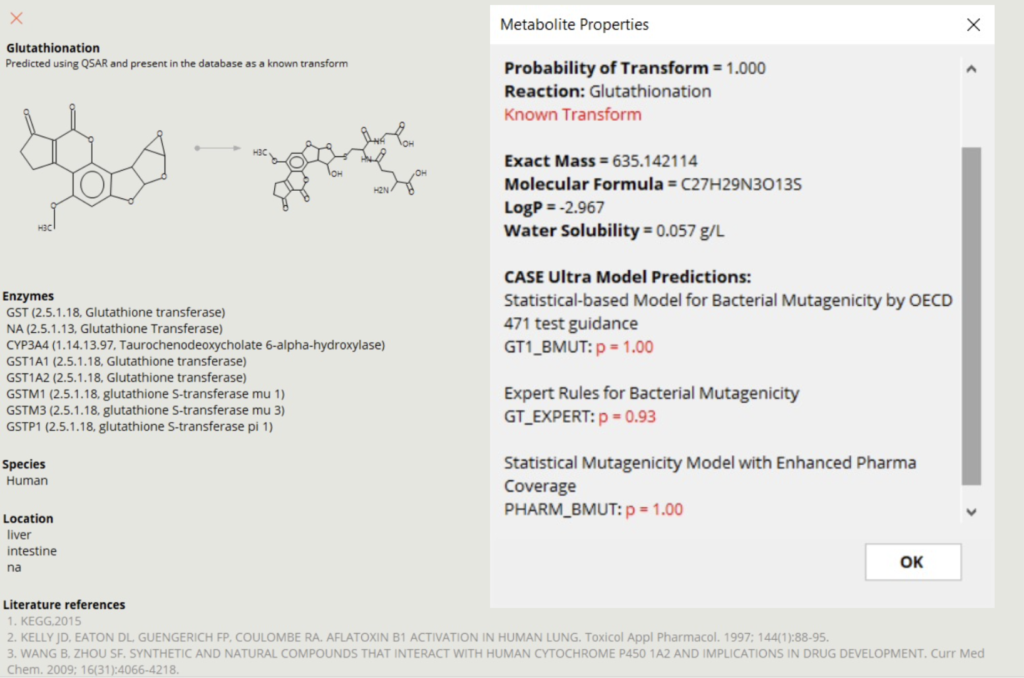
META Ultra contains data for experimentally observed metabolites of drugs and chemicals. The database is composed of 14,697 of known human metabolites and their parent compounds, 17,488 known human metabolic transformations, 4511 responsible enzymes located in 10 organs, 329 types of reactions and yields, and so on. A collection of 36,121 unique literature references supports these experimental findings. Experimentally observed metabolites are indicated by the program.

Each metabolite contains information including the reaction type, enzymes involved, species, location, and literature references. Molecular properties are calculated for logP and water solubility. META Ultra can also identify reactive metabolites by connecting to CASE Ultra.

License packages are customized to meet the needs of the user. Factors that affect pricing include length of license, number of endpoints, and number of users. Please contact us to request a quote.